Welcome To Developmental Biology Group
For decades, developmental biologists have been fascinated with one question: “How can a fertilized egg give rise to a complex multicellular organism?” This process requires cells in the developing embryo to divide, migrate, and differentiate in a coordinated fashion to create a functional organism. We at ARI are focused on understanding how the behaviors of cells in the developing embryo direct distinct processes and how genes contribute to the overall development. We are using a diverse group of model organisms and paradigms to address some of the fundamental questions in animal development. These include Hydra, Drosophila, zebrafish, and the chick embryo.
Topics under investigation
Some of the interesting topics that are currently under intense investigation are: Cell-cell communication, molecular basis of pattern formation and stress response, synapse formation at the neuromuscular junction, regulation of autophagy during development, heart development and regeneration, blood vessel development, and stem cell biology. We are currently developing TALEN and CRISPR-Cas-based genome editing techniques to be used for generating allele-specific mutant organisms. Importantly, we foster collaborative research and nurture relationships with multidisciplinary organizations from within and outside India.

Name: Dr. Anuradha Ratnaparkhi
Designation: Scientist – F
Brief Background :
Contact Details :
020-25325043/44 (Office)
Educational Qualification
- BSc (Zoology), M.S. University, Vadodara.
- MSc (Biotechnology), S.P.Pune University, Pune.
- PhD, NCBS-TIFR, Bangalore.
- Post-doctoral training: California Institute of Technology (Caltech), Pasadena, USA. University of California, Los Angeles (UCLA), Los Angeles, USA.
Research Interests
In our lab, we study signalling mechanisms in the nervous system that regulate the assembly of synapses, their function, and systemic communication during organ growth and maturation. We are particularly interested in the regulation of these physiological processes by the Mon1-CCZ1 (MC1) complex that regulates cellular trafficking along the endo-lysosomal pathway- a pathway that is impaired in neurodegenerative diseases such as Charcot-Marie-Tooth 2B.
The Drosophila larval neuromuscular junction (NMJ) is a well-established model for studying synaptic development and function. Similar to the mammalian central nervous system, these synapses are glutamatergic. Studies have shown that post-synaptic glutamate receptors play a key role in synaptic homeostasis. A decrease in receptor levels triggers a retrograde signal (muscle to nerve) that eventually leads to a compensatory increase in neurotransmitter release, thus maintaining homeostasis.
In our lab, we have identified Mon1 as a transsynaptic factor that negatively regulates glutamate receptor levels in an anterograde manner at the NMJ. In addition, Mon1 also functions as an important molecular component of a neural circuit that controls gonad maturation through the regulation of insulin-like peptides. Our current investigations are aimed at understanding how modulation of endo-lysosomal trafficking events regulates these physiological events.
Another area of study in the lab involves identifying mechanisms that regulate glial morphology and its implications for neuronal function. Of particular interest is the GPCR signalling pathway activated by the ligand Folded gastrulation (Fog). Ongoing studies are aimed at identifying cell-autonomous and cell non-autonomous factors that modulate this pathway to regulate glial morphogenesis.
Publications:
- Lakshmi Bugga, Anuradha Ratnaparkhi, and Kai Zinn (2009). The cell surface receptor Tartan is a potential in vivo substrate for the receptor tyrosine phosphatase Ptp52F. Mol Cell Biol. 29(12):3390-400.
- Ratnaparkhi A. (2013) Signaling by Folded gastrulation is modulated by mitochondrial fusion and fission. J Cell Sci. 1;126:5369-76.
- Deivasigamani S, Verma HK, Ueda R, Ratnaparkhi A, Ratnaparkhi GS. .(2014). A genetic screen identifies Tor as an interactor of VAPB in a Drosophila model of amyotrophic lateral sclerosis. Biol Open. 3(11):1127-38
- Deivasigamani S, Basargekar A, Shweta K, Sonavane P, Ratnaparkhi GS, Ratnaparkhi A (2015).A Presynaptic Regulatory System Acts Transsynaptically via Mon1 to Regulate Glutamate Receptor Levels in Drosophila. Genetics. 2015 Oct;201(2):651-64.
- Dhiman N, Shweta K, Tendulkar S, Deshpande G, Ratnaparkhi GS, Ratnaparkhi A (2019) Drosophila Mon1 constitutes a novel node in the brain-gonad axis that is essential for female germline maturation. Development, 146(13):dev166504.
- Rohit H KrishnanA, Shweta Tendulkar, Senthilkumar Deivasigamani , Anuradha Ratnaparkhi, Girish S Ratnaparkhi (2019). Monensin sensitive 1 regulates dendritic arborization in Drosophila by modulating endocytic flux. Frontiers in Cell and Developmental Biology, 2;7:145.
- Anagha Basargekar, Shweta Yogi, Zeeshan Mushtaq, Senthilkumar Deivasigamani, Vimlesh Kumar, Girish Ratnaparkhi, Anuradha Ratnaparkhi (2020). Drosophila Mon1 and Rab7 interact to regulate glutamate receptor levels at the neuromuscular junction. Int. J.Dev.Biol, 64(4-5-6):289-297.
- Kumari Shweta, Anagha Basargekar and Anuradha Ratnaparkhi (2021) FGFR/Heartless and Smog interact synergistically to negatively regulate Fog mediated G-protein coupled receptor signaling in the Drosophila nervous system. G3 (Bethesda), 11(3), jkaa029.
- Tendulkar S, Hegde S, Garg L, Thulasidharan A, Kaduskar B, Ratnaparkhi A, Ratnaparkhi GS (2022). Caspar, an adapter for VAPB and TER94, modulates the progression of ALS8 by regulating IMD/NFκB-mediated glial inflammation in a Drosophila model of human disease. Hum Mol Genet. 231(17):2857-2875
- Ratnaparkhi A, Sudhakaran J (2022). Neural pathways in nutrient sensing and insulin signalling. Front Physiol.13:1002183
- Rai P, Ratnaparkhi A, Kumar Roy J (2023). Rab11 rescues muscle degeneration and synaptic morphology in the park13/+ Parkinson model of Drosophila melanogaster. Brain Res. 1816:148442.
Book Chapter/Book Review:
- Kriti Chaplot, Anuradha Ratnaparkhi, Girish Ratnaparkhi (2019). ‘Understanding motor disorders using flies’in M. Mutsuddi and A. Mukherjee (Eds), ‘Insights into Human neurodegeneration: Lessons learnt from Drosophila’, 131-162 (Springer).
- Anuradha Ratnaparkhi (2020). ‘Introduction to the embryonic nervous system in Drosophila’ in S.C.Lakhotia and H.A. Ranganath (Eds), ‘ Experiments with Drosophila for Biology Courses’, Indian Academy of Sciences, Bengaluru.
- Ratnaparkhi, A (2021). Code of Life: Revolutions in Genetics. Mohan Sundara Rajan. Current Science, 120: 1256.

Google Scholar / Social Media
Name: Dr. Chinmoy Patra
Designation: Scientist – E
Brief Background :
Dr. Chinmoy Patra has pursued his undergraduate studies in Pharmaceutical Science from Jadavpur University, Kolkata. He secured All India Rank-1 in Graduate Aptitude Test in Engineering (GATE) – 2005 in Pharmaceutical Science. He has received his Master’s degree (M.Tech) fromIIT Kharagpur and was awarded DAAD Fellowship(2006) tocomplete his M.Tech project atTU Dresden, Germany.
He has pursued his doctoral work under the guidance of Dr. Felix B. Engle atthe Max-Planck-Institute forHeart and Lung Research, Germany. His doctoral work leads to the understanding the role of extracellular matrix molecule nephronectin in cardiac valve development in zebrafish. Subsequently, he worked with Prof. Didier Stanier as a postdoctoral fellow at the MPI for heart and Lung Research, Germany. His research works lead toa number ofpublications in reputed peer reviewed journals like Development, Cardiovascular research, Biomaterials, PNAS USA, Blood, Journal of Neuroscience etc.
In 2014, Dr. Patra has joined Agharkar Research Institute,Pune and established hislab with the research focus to understand the cellular and molecular regulation behind the cardiovascular development, tissue homeostasis, and heart regeneration using zebrafish as a model organism.
His group has been published in Science, Cell & Bioscience, Thrombosis and Haemostasis, Scientific reports etc. and research work has also been discussed in several news media like The Indian Express, Research Mattersand the Times of India. He has received several prestigious awards like, Max-Planck Partner Group Award, in 2015, Early Carrier Research Award from Department of Science and Technology, New Delhi in 2016, DBT/Wellcome Trust India Alliance Intermediate Fellowship in 2019.
Contact Details :
020-2532-5046 (Telephone)
7743875812 (Mobile)
Educational Qualification
- From 2018 : Scientist D at the Agharkar Research Institute, Division of Developmental Biology, Pune, INDIA
- 2014-2018: Scientist C at the Agharkar Research Institute, Division of Developmental Biology, Pune, INDIA
- 2012-2014: Post Doctoral fellow with Prof. Didier Stainier. Department of Developmental Genetics, Max-Planck-Institute for Heart and Lung Research, Germany.
- 2010-2012 : Post Doctoral fellow with Prof. Felix B. Engel. Department of Heart Development and Regeneration, MPI for Heart and Lung Research, Germany.
- 2007-2010 : Performed Doctoral thesis work under the guidance of Dr. F.B. Engel at Max Planck Institute for Heart and Lung Research, Germany
Thesis title: Nephronectin regulates cardiac valvedevelopment via BMP4-HAS2 signaling in zebrafish. (PhD Degree Received-November, 2011) Grade-Very good
Research Interests
Development of an organ or new life is a spectacular process, which is controlled by spatiotemporal gene expression. A considerable part of any tissue is extracellular space, which is filled up with highly complex networks of macromolecules like proteins, proteoglycans etc. as well as relatively simpler molecules such as saccharides and water. Historically the importance of the extracellular matrix (ECM) was as an inert supporting structure to maintain the tissue architecture. However, recent studies from my and other laboratories clearly indicate that ECM components also play major roles during organogenesis, tissue homeostasis, in organ function, in disease, and in regeneration by regulating signalling networks. Thus we can say ECM is the hub of master regulators.
My lab is interested to understand how ECM molecules and adhesion G-protein coupled receptors (aGPCRs) are involved in organ development, tissue homeostasis, organ function and regeneration at the cellular level, using zebrafish as model organism. We investigate our questions utilizing genetic (transgenesis, genome editing with the help of state-of-the-art technology TALEN and CRISPR-Cas), and pharmacological approaches. One of our goals is to understand the crosstalk between different cell types via ECM components during heart development and heart regeneration.
We are looking for enthusiastic, highly motivated students wish to work in the field of cardiovascular biology and genome editing. We also offer scientific training to Masters and Bachelor students.
Key Publications:
- Barrodia P, Patra C and Swain RK. EF-hand domain containing 2 (Efhc2) is crucial for distal segmentation of pronephros in zebrafish. Cell & Bioscience (ISSN#2045-3701) 2018; 8:53.
- Patra C*, Kontarakis Z, Kaur H, Rayrikar A, Mukherjee D and Stainier DYR. The zebrafish ventricle: A hub of cardiac endothelial cells for in vitro cell behavior studies. Scientific Reports (ISSN#2045-2322) 2017; 7:2687.
* Corresponding Author
- Mokalled MH, Patra C, Dickson AL, Endo T, Stainier DYR and Poss KD. Injury-induced ctgfa directs glial bridging and spinal cord regeneration in zebrafish. Science (ISSN#1095-9203)2016; 354 (6312):630-634.
- Patra C,Boccaccini AR and Engel FB. Vascularization for cardiac tissue engineering: the extracellular matrix. Thrombosis and Haemostasis (ISSN#1538-7836) 2015; 113(3):532-47.
- Patra C, van Amerongen MJ, Ghosh S, Ricciardi F, Sajjad A, Novoyatleva T, Mogha A, Monk KR, Mühlfeld C and Engel FB. Organ-specific function of adhesion G protein-coupled receptor GPR126 is domain dependent. Proc. Natl. Acad. Sci. USA(ISSN#1091-6490)2013; 110 (42):16898-16903.
- Mogha A, Benesh AE, Patra C, Engel FB, Schöneberg T, Liebscher I, Monk KR. Gpr126 Functions in Schwann Cells to Control Differentiation and Myelination via G-Protein Activation. J Neurosci.(ISSN 0270-6474)2013; 33(46):17976-17985.
- Patra C,Ricciardi F, and Engel FB.The functional properties of nephronectin: an adhesion molecule for cardiac tissue engineering. Biomaterials (ISSN#0142-9612)2012;33(17):4327–4335.
- Patra C, Talukdar S, Novoyatleva T, Reddy S,Mühlfeld C, Kundu B, Kundu SC and Engel FB. Silk protein fibroin from Antheraeamylitta for cardiac tissue engineering. Biomaterials (ISSN#0142-9612)2012;33(9):2673-2680.
- Patra C, Diehl F, Ferrazzi F, van Amerongen MJ, Novoyatleva T, Schaefer L., Mühlfeld C, Jungblut B and Engel FB. Nephronectin regulates atrioventricular canal differentiation via Bmp4-Has2 signaling in zebrafish. Development (ISSN#0950-1991) 2011; 138(20):4499-4509.

Google Scholar / Social Media
Name: Dr. Bhupendra V. Shravage
Designation: Scientist – E
Brief Background :
Dr. Bhupendra V. Shravage completed his doctoral studies under the guidance of Prof. Dr. Siegfried Roth at the Institute for Developmental Biology, University of Cologne, Germany. He investigated the role of “TGF-beta and EGF signaling pathways involved in patterning of dorsal chorion structures in Drosophila”.
He moved to University of Pittsburgh, USA, to work with Dr. Beth Roman as a post-doctoral fellow on understanding the role of Alk1 signaling in blood vessel development during zebrafish embryogenesis. He did his second post-doctoral with Prof. Eric Baehrecke at UMass Medical School,USA, on “Autophagy in cancer and cell death”.
During his tenure at UMass, they discovered the multi-functional role of Atg6 (Beclin1 in mammals) in various membrane-trafficking pathways and haematopoiesis in Drosophila. Since 2014 Dr. B. V. Shravage is working as a Scientist at the Agharkar Research Institute, Pune, India and is affiliated to Savitribai Phule Pune University, Pune, India.
His research work is currently focused on understanding molecular regulation and function of autophagy in the germline. He is recipient of the prestigious Ramalingaswami Fellowship (Dept of Biotechnology, Govt. of India) and Early Career Research Award (Dept of Science and Technology)
Contact Details :
020-25325048, 020-25325045, 020-25325076 (Telephone)
Educational Qualification
- BSc Microbiology Abasaheb Garware College, University of Pune, 1998.
- MSc Marine Biotechnology, Goa University, 2000
- Dr. rer. Nat (German PhD), University of Cologne, Germany 2006.
Research Interests
Macro-autophagy(termed as autophagy henceforth) is a conserved process that facilitates degradation of cytoplasmic components including toxic proteins/aggregates and damaged organelles via the lysosome. Autophagy functions have been identified from gametogenesis to maintenance of cellular homeostasis during aging.
We are interested in understanding the role of autophagy in animal development and in diseases such as cancer and neurodegeneration. We use Drosophila melanogaster, as a model system to study autophagy in various cell typesand in distinct cellular contexts. Currently, we are focused on unravelling novel functions of autophagy and its regulation in stem cellsincluding germline stem cells.
Key Publications:
- Klionsky D.,Shravage B.V., et al.,(2020)Guidelines for the Use and Interpretation of Assays for Monitoring Autophagy (4th edition)Autophagy In press
- Nilangekar, K. S., Murmu N., Sahu G. and Shravage B.V. (2019) Generation and Characterization of Germline-Specific Autophagy and Mitochondrial Reactive Oxygen Species Reporters in Drosophila. Front Cell Dev Biol. 2019 Apr 3;7:47
- Nilangekar K.S., Shravage B.V. (2018) Mitochondrial Redox Sensor for Drosophila Female Germline Stem Cells. In: Turksen K. (eds) Autophagy in Differentiation and Tissue Maintenance. Methods in Molecular Biology, vol 1854. Humana Press, New York, NY.Reserch highlighted on the cover of the book series.
- Bali, A. and Shravage, B. V. (2017). Characterization of the autophagy related gene-8a (Atg8a) promoter in drosophila melanogaster. J. Dev. Biol.61, 551–555.
- Dixit, N. S., Shravage, B. V. and Ghaskadbi, S. (2017). Identification and characterization of the autophagy-related genes Atg12 and Atg5 in hydra. J. Dev. Biol.61, 389–395.Research highlighted on the cover of the journal.
- Chang, T.-K., Shravage, B. V., Hayes, S. D., Powers, C. M., Simin, R. T., Wade Harper, J. and Baehrecke, E. H. (2013). Uba1 functions in Atg7- and Atg3-independent autophagy. Cell Biol.15, 1067–78.
- Shravage, B. V., Hill, J. H., Powers, C. M., Wu, L. and Baehrecke, E. H. (2013). Atg6 is required for multiple vesicle trafficking pathways and hematopoiesis in Drosophila. Development140, 1321–9.

Google Scholar / Social Media
Name: Dr. Arindam Bhattacharjee
Designation: SERB Ramanujan Fellow
Brief Background :
Dr. Arindam Bhattacharjee pursued his undergraduate studies at the University of Calcutta. In 2012, he joined Dr. Kalyan Mitra’s lab at CSIR-Central Drug Research Institute, Lucknow to perform doctoral work on drug-induced cell death mechanisms,which turned out to be regulated by autophagy. He then extensively trained in high-resolution microscopy techniques (e.g., TEM) to study the autophagic process on an ultrastructural level.
This, coupled with a well-timed 2016 Nobel to Prof. Yoshinori Ohsumi, motivated him to pursue a postdoc in the field of autophagy in the lab of Prof. Gabor Juhaszat the Biological Research Centre, Hungary in 2017. Here, as an NKFIH OTKA Postdoctoral Fellow, he worked on Drosophila models of selective ubiquitin autophagy, SNARE proteins, and lysosomal ion channels. He returned to India in 2024 with a Ramanujan Fellowship from SERB and a Ramalingaswami Fellowship from DBT.
Since 2024, Dr. Bhattacharjee has joined Agharkar Research Institute to work on understanding lysosomal diversity on a physiological level. Specifically, his group will try to establish a multi-dimensional model of the intracellular population of lysosomes using quantitative microscopy-based approaches and then challenge cells to stressors such as proteotoxicity (e.g., mutant Huntingtin expression), lysosomal ionic imbalance, and infection to study the shift in lysosomal behavior under those setups. Since lysosomes are the endpoint of multiple cellular pathways and often implicated in diseases, the findings will ultimately be used to design novel interventions to target lysosomal catabolism. Scientific training opportunities for interested BSc and MSc studentsareopen.
Contact Details :
020-2532-5155 (Telephone)
Educational Qualification
- 2024-present: SERB Ramanujan Fellow (Scientist-D equivalent) at the Agharkar Research Institute, Division of Developmental Biology, Pune, India.
- 2020-2023: NKFIH OTKA Postdoctoral Fellow, Institute of Genetics, Biological Research Centre, Szeged, Hungary
- 2017-2020: Postdoctoral research associate, Institute of Genetics, Biological Research Centre, Szeged, Hungary.
- 2018: PhD, Jawaharlal Nehru University, New Delhi, India.
- 2012-2017: CSIR JRF & SRF at CSIR-Central Drug Research Institute, Lucknow, India.
- 2011: MSc, Asutosh College, University of Calcutta (Ranked Ist within batch).
- 2009: BSc, BKC College, University of Calcutta (Ranked Istwithin batch).
Research Interests
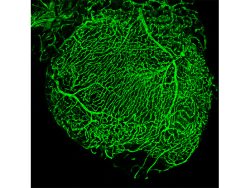
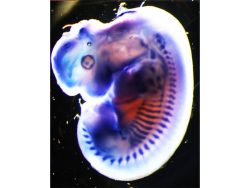
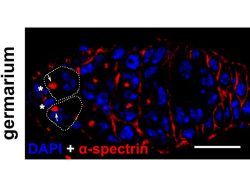
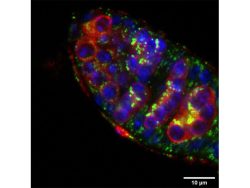
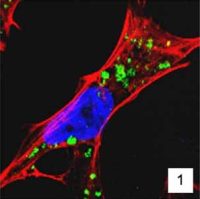
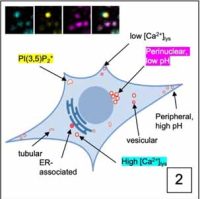
Our interest in the physiology of lysosomes [green dots in Panel 1] stems from a few observations: firstly, they are the endpoint of multiple degradative pathways including endocytosis, autophagy, and phagocytosis, and therefore, their function is critically important to cellular health. Second, lysosomes offer remarkable plasticity, regulated through a series of biogenesis, repair, and removal steps (Trends Cell Biol. 2022;32(7):597-610). Third, they are uniquely acidic (pH 4.5-5), although not all lysosomes are equally so (J Cell Biol. 2016;212(6):677-692).This gives rise to a diverse array of highly dynamic organelles responding to cellular cues [Panel 2], which have thus far been poorly characterized as a whole.
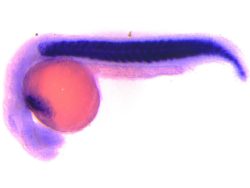
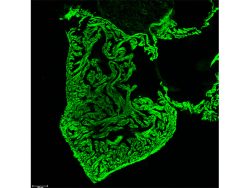
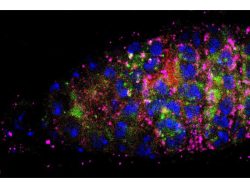
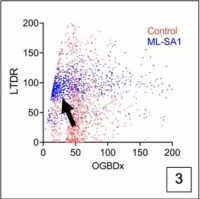
Using genetic and microscopic tools, we will aim to build a model of intracellular lysosomal diversity taking into account its physical and biochemical features and study its response under scenarios such as amino acid starvation (autophagy), lysosomal calcium release, lysosomal membrane damage, proteotoxicity (expression of toxic and non-toxic cargoes such as p62 and mHtt), and infection [Panel 3 shows an example of alysosomal response upon opening an ion channel in HEK-293 cells, where the individual vesicles shift from high luminal Ca2+ to low luminal Ca2+ (x-axis), and from high luminal pH to low luminal pH (y-axis; increase in fluorescence corresponds to acidification)]. The ultimate goal would be to use genetic/pharmacological means to manipulate lysosome behavior during stress toward fulfilling their catabolic goals.
Key Publications:
- Bhattacharjee A*, Abuammar H*, Juhász G. Lysosomal activity depends on TRPML1-mediated Ca2+ release coupled to incoming vesicle fusions. J Biol Chem 2024; 300(12):107911.
- Laczkó-Dobos H*, Bhattacharjee A*, Maddali AK, Kincses A, Abuammar H, Sebők-Nagy K, Páli T, Dér A, Hegedűs T, Csordás G, Juhász G. PtdIns4P is required for the autophagosomal recruitment of STX17 (syntaxin 17) to promote lysosomal fusion. Autophagy. 2024; Mar 8:20(7):1639-1650.
- Bhattacharjee A*, Ürmösi A*, Jipa A, Kovács L, Deák P, Szabó Á, Juhász G. Loss of ubiquitinated protein autophagy is compensated by persistent cnc/NFE2L2/Nrf2 antioxidant responses. Autophagy. 2022;18(10):2385-2396.
- Bhattacharjee A*, Hasanain M*, Kathuria M, Singh A, Datta D, Sarkar J, Mitra K. Ormeloxifene-induced unfolded protein response contributes to autophagy-associated apoptosis via disruption of Akt/mTOR and activation of JNK. Sci Rep. 2018;8(1):2303.
- Hasanain M*, Bhattacharjee A*, Pandey P, Ashraf R, Singh N, Sharma S, Vishwakarma AL, Datta D, Mitra K, Sarkar J. α-Solanine induces ROS-mediated autophagy through activation of endoplasmic reticulum stress and inhibition of Akt/mTOR pathway. Cell Death Dis. 2015;6(8):e1860. (*=equal contribution)
Reviews:
- Abuammar H, Bhattacharjee A, Simon-Vecsei Z, Blastyák A, Csordás G, Páli T, Juhász G. Ion Channels and Pumps in Autophagy: A Reciprocal Relationship. Cells. 2021 Dec 14;10(12).
- Bhattacharjee A*, Szabó Á*, Csizmadia T, Laczkó-Dobos H, Juhász G. Understanding the importance of autophagy in human diseases using Drosophila. J Genet Genomics. 2019 Apr 20;46(4):157-169.
Patents:
- Goel A, Sharma A, Mitra K, Bhattacharjee A, Kathuria M. Substituted fluoroanthene-7- carbonitriles/esters as fluorescent dyes for cell imaging applications. WO2014147642A1.

Name: Ms. Rohini Jayant Londhe
Designation: Technical Officer – B
Job Description :
- Maintenance of an important model system: Hydra And providing the start-up culture kits for the same all over India.
- Providing Technical support to the scientists and researchers in the group.
- Operating the central Facility of ARI i.e., Confocal Microscope System for all the researchers at ARI
Contact Details :
020 25325047, 9623994662 (Telephone)
Educational Qualification
- B.Sc. Zoology
- M.Sc. Zoology (Specialization: Developmental Biology)
Publications/ Achievements/ Awards
Publications:
- Description and phylogenetic characterization of Hydra strain from Naukuchiatal, (Uttarakhand, India) and comparison with other Hydra strains. Current Science, Vol. 113, No. 9, 10 November 2017.
- UV induced foot duplication in regenerating hydra is mediated by metalloproteinases and modulation of the Wnt pathway. Int. J. Dev. Biol, 60:111-117, April 2016.
Award:
- Professor Sukhatme Award for Best Performance on 26th January 2016 (For the year 2015-16)

Name: Mr. Mahadeo Daware
Designation: Technical Officer – B
Job Description :
Breeding, care, management and experimentationson laboratory animals, Confocal imaging of the biological and other samples, Guidance of use and maintenance of laboratory equipment’s. Preparations of SOP/Protocols of routine laboratory work andsupervision etc.
Contact Details :
020 02025325047 (Telephone)
9922288705 (Telephone)
Educational Qualification
- B.Sc. Zoology
- Zoology
Publications/ Achievements/ Awards
Publications:
- Reproductive toxicity of Piperine in Swiss albino mice. Planta Medica, 2000; 66(3):231-6. doi: 10.1055/s-2000-8560.रिप्रोडक्टिव्हटॉक्सिसिटीऑफपायपेरीनइनस्विसअल्बिनोमाइस.प्लानटामेडिका, 2000; 66(3):231-6. doi: 10.1055/s-2000-8560

Name : Amey Rayrikar
Designation : Ph.D. (SRF)
Brief Background :
Amey Rayrikar has completed Masters in Biotechnology from Pune University. He cleared BET-DBT JRFin 2015. He has joined as a Junior Research Fellow in Agharkar Research Institute and currently pursuing his Ph.D.
Contact Details :
Educational Qualification
- B.Sc. Biotechnology, 2012
- M.Sc. Biotechnology, 2014
Research Interests
He’s interested in understanding the cellular and molecular eventsin vertebrate’s development. His doctoral work focuses onunderstanding the cellular and molecular events those take place during intervertebral tissue morphogenesis.
Publications / Patents / Varieties
Patra C*, Kontarakis Z, Kaur H, Rayrikar A, Mukherjee D and Stainier DYR. The zebrafish ventricle: A hub of cardiac endothelial cells for in vitro cell behavior studies. Scientific Reports (ISSN#2045-2322) 2017; 7:2687

Name : Ms. Bhagyashri Joshi
Designation : Ph.D – SRF
Brief Background :
I have completed my bachelors and masters in biochemistry and currently pursuing PhD in biotechnology.
Contact Details :
Educational Qualification
- B.Sc (Biochemistry) 2012
- M.Sc (Biochemistry) 2014
Research Interests
Research Topic : The role of CELSR1 in morphogenesis using zebrafish as a model organism.
Investigating the role of CELSR1 in morphogenesis using zebrafish as a model organism.
Cadherin EGF LAG seven-pass G-type receptors 1, 2 and 3 (CELSR1-3) receptor protein family, a sub-class of adhesion G-protein coupled receptor has wide spectrum of functions in development and maintenance. They have characteristic long N-terminal fragment, seven transmembrane and c terminal fragment. I am interested in studying the expression patternand role of CELSR family in development using genome editing and pharmacology techniques.
Publications / Patents / Varieties
Name : Mr. Ganesh Wagh
Designation : Junior research Fellow
Brief Background :
Mr. Ganesh Wagh hascompleted his Bachelor of science in Biotechnology from COCSIT Latur andMasters of science in Biotechnology from SRTMUN University, Nanded in 2015 and qualified CSIR-JRF in 2016.
Currently working as a Ph.D scholar at MACS-Agharkar research institute, Pune 411004
Contact Details :
Educational Qualification
- B. Sc Biotechnology 2013
- M. Sc. Biotechnology 2015
Research Interests
Research Topic : Elucidation of the role of selected secreted molecules in zebrafish development.
- Understanding the role of extracellular matrix molecule like sparc, mif,in organ development and heart regeneration.
Publications / Patents / Varieties

Name : Karan Selarka
Designation : U.G.C. JRF (Ph.D Student)
Brief Background :
I am a Biochemistry Master’s from Fergusson College, Pune and was a Project JRF at ICMR- National Institute of Virology, Pune working on “Metagenomics Project.”I have always dreamt of becoming a researcher and hence I registered for Ph.D.
Qualified:
- Joint CSIR-UGC-NET Life Sci. (Dec2019): AIR 72.
- GATE2020: Life Sciences: AIR 125.
- TIFR-JGEEBILS (GS2020).
- GATE2019: Life Sciences: AIR 578
Contact Details :
Educational Qualification
- B.Sc. Chemistry, Botany, Zoology (SGBAU, Amravati)
- M.Sc. Biochemistry (SPPU, Pune)
Research Interests
Autophagy is a degradative mechanistic process, important for cell’s maintenance and survival throughout its lifespan. Autophagy balances cellular turnover, and this degradative process marks the beginning of synthesis. It has been well understood that, disruption/ obstruction in autophagy eventually leads to diseases/ disorders and hence it becomes important to understand the process which may lead to eradication of the diseases/ disorders. Therefore, it is important to understand how autophagy is regulated and are there any ‘still unknown (novel) regulators’ of autophagy, which may be used as tools to act with/ on for prevention/ cure of disorders.
Publications / Patents / Varieties

Name : Kiran Nilangekar
Designation : PhD Student
Brief Background :
After the post graduate degree in Virology, I was a Project Fellow at the Cytoskeleton and Cell shape lab at IISER, Pune after which I joined Dr. Bhupendra Shravage’s lab as Junior Research Fellow before starting my PhD here. It has been a nice experience through the different biological systems.
Contact Details :
Educational Qualification
- B.Sc. Microbiology
- M.Sc. Virology
Research Interests
Autophagy is a cellular process that keeps cellular damage in check by its sequestration and degradation. Many such homeostatic processes respond to dysregulation which occurs during aging. Autophagy in the context of stem cells is understudied. Stem cells exist in their undifferentiated state or “stemness” because of a microenvironment which is called the “niche”. Niche comprises of the physical confinement, accessory cells and an interplay of cellular signalling molecules.
I am interested in the role of autophagy for stem cells maintenance. Apart from accumulation of damage, reactive oxygen species (ROS) is another important hallmark of aging. We have generated and characterized germline specific reporters for mitochondrial reactive oxygen species and will employ them to study ROS during aging. Our approach for the study is primarily by microscopy. Although studying a complicated process like autophagy in highly specialized cells like niche and stem cells is challenging, we hope that with this study we add to the knowledge about autophagy, stem cells, their niche and this would also serve as a model for similar studies.
Publications / Patents / Varieties
- Nilangekar, K., Murmu, N., Sahu, G., &Shravage, B. V. (2019). Generation and characterization of germline-specific autophagy and mitochondrial reactive oxygen species reporters in Drosophila. Frontiers in Cell and Developmental Biology, 7(APR), 1–18. https://doi.org/10.3389/fcell.2019.00047
- Nilangekar, K. S., &Shravage, B. V. (2019). Mitochondrial Redox Sensor for Drosophila Female Germline Stem Cells. Methods in Molecular Biology (Clifton, N.J.), 1854, 13–20. https://doi.org/10.1007/7651_2018_167

Name : Minal Ayachit
Designation : Junior research Fellow
Brief Background :
After completing my graduation and post-graduation studies in Biotechnology, I was interested in working in a research lab to understand mechanisms of cell biology. The resourceful atmosphere and well equipped lab under the guidance of Dr. Shravage was conducive for my interests.
Contact Details :
Educational Qualification
- B.Sc. in Biotechnology
- M.Sc. in Biotechnology
Research Interests
Autophagy is a process where constituents of the cell such as proteins and organelles that are damaged or not required anymoreare sequestered within a double membrane structure called the Autophagosome, which then fuses with the lysosome.
The lysosomal enzymes degrade the cargo and recycle the raw materials. It is a cytoprotective mechanism that increases the lifespan of the cell by maintaining homeostasis.
My research interest includes understanding the role of autophagy and specifically, mitophagy i.e degradation of the damaged mitochondria, in the maintenance of germline stem cellsthat reside within the Drosophila melanogaster ovary.
Publications / Patents / Varieties

Name: Nidhi Murmu
Designation: Ph.D. Student
Brief Background :
After completing M.Sc. in Biotechnology, I joined Dr.Shravage’s lab as a Ph.D. student in developmental biology group.
Contact Details :
Educational Qualification
- B.Sc. Biotechnology
- M.Sc. Biotechnology
Research Interests
Autophagy is a stress response process where protein aggregates and damaged organelles are degraded via delivery to lysosomes. The dysregulation of this proteostatic process increases with age which ultimately leads to disruption of cellular homeostasis.
I am interested in understanding the role of autophagy in germline stem cell (GSC) aging and how it is regulated in Drosophila ovary.
List of Publications/Patents/Varieties
- Nilangekar, K., Murmu, N., Sahu, G. and Shravage, B.V., 2019. Generation and characterization of germline-specific autophagy and mitochondrial reactive oxygen species reporters in Drosophila. Frontiers in cell and developmental biology, 7, p.47.

Name: Ms. Ashwini Punde
Designation: DBT-JRF
Brief Background :
I have completed my Master’s in Biotechnology from SPPU, Pune.
Contact Details :
Educational Qualification
Research Interests
Research Topic: Exploration of the role of Formin Homology 2 Domain Containing 3 (FHOD3) in cardiac maintenance.
My research interest focuses on investigating the role of actin-binding proteins in cardiac development as well as their involvement in cardiomyopathy using zebrafish as a model.
List of Publications/Patents/Varieties

Name: Ms. Shweta Kalke
Designation: UGC-JRF
Brief Background :
I have completed my Master’s in Zoology from Ramnarain Ruia Autonomous College, Mumbai.
Contact Details :
Educational Qualification
Research Interests
Research Topic: Identifying the cardiac pro-regenerative ECM molecules and their cellular and molecular function.
Identifying the role of extracellular matrix (ECM) molecules in Heart Regeneration using zebrafish as a model.
List of Publications/Patents/Varieties

Name: Mrunmayee Kulkarni
Designation: Ph.D. Student (CSIR-JRF)
Brief Background :
I completed my masters and bachelors in Biotechnology from Abasaheb Garware college, Pune. After clearing my CSIR, I joined PhD under the guidance of Dr. Bhupendra Shravage, Developmental Biology Group.
Contact Details :
Educational Qualification
Research Interests
My work is based on investigating the underlying mechanisms of mitochondrial homeostasis specifically during the process of oogenesis of Drosophila melanogaster. I am currently investigating the role of Dpp/BMP pathway in regulation of mitochondrial homeostasis and mitophagy in Drosophila oogenesis.
List of Publications/Patents/Varieties

Name: Mr. Swarnav Bhakta
Designation:
Brief Background :
Contact Details :
Educational Qualification
Research Interests
Research Topic: Role of actin-binding protein in heart development and maintenance.
List of Publications/Patents/Varieties

Name: Ms. Shreya Maity
Designation: Research Fellow
Brief Background :
Contact Details :
Educational Qualification
Research Interests
Research Topic: Exploration of the cellular and molecular mechanism underlying the development of cardiomyopathy.
List of Publications/Patents/Varieties
- Patra C,*Rayrikar A, Wagh G, Kleefeldt F,Roshanbinfar K, Cop F,Nikolic I,Schmidt MHH, Acker-Palmer A, Ergün S, Engel FB*. Nephronectin is Required for Vascularization in Zebrafish and Sufficient to Promote Mammalian Vessel‐Like Structures in Hydrogels for Tissue Engineering. J. of the American Heart Association, (ISSN#2047-9980), 2025, 14, https://doi.org/10.1161/JAHA.123.037943.
- Chakraborty G, K. Ahire, Joshi B, Patra C*.Tri-n-butyl phosphate inhibits neurogenesis and motor functions during embryonic development in zebrafish. Aquatic Toxicology, (ISSN#1879-1514) 2025, 279, 107203.
- Manikandan M, Chhatar S, Dey S, Panda TR, Chakraborty S, Ray P, Patra C*, Patra M*. Analysis of Antiangiogenic Potential and Cell Death Mechanism of a Kinetically Inert Platinum Antitumor Agent. Medicinal Chemistry Letters.(ISSN#1948-5875) 2024, 15, 1482-1490.
- Gadre S, Manikandan M, Chakraborty G, Rayrikar A, Paul S, Patra C*, Patra M*. Development of A Highly In Vivo Efficacious Dual Antitumor and Antiangiogenic Organoiridium Complex as Potential Anti-Lung Cancer Agent. J Med Chem.(ISSN#0022-2623) 2023, 66, 13481-13500.
- Vaidya SP, Manikandan M, Chhatar S, Dey S, Patra C, Patra M*. A Hydrolytically Stable Oxo-Rhenium(V) Antitumor Agent for Synergistic Combination Therapy with Cisplatin: From Synthesis and Mechanistic Studies to Toxicity Assessment in Zebrafish. Chem. Front.(ISSN#2052-1553) 2023, 22, September.
- Takahi M, Taira R, Onozuka J, Sunamura H, Kondow A, Nakade K, Nakashima K, Sato I, Hayashi Y, Patra C, Ohnuma K. Xenograft of human pluripotent stem cell-derived cardiac lineage cells on zebrafish embryo heart. BiochemBiophys Res Commun.(ISSN#1090-2104)2023; 14;S0006-291X(23)00795-7.
- Rayrikar A, Wagh G, Santra MK, and Patra C*.Ccn2a-FGFR1-SHH signaling is necessary for intervertebral disc homeostasis and regeneration in adult zebrafish. Development(ISSN#0950-1991) 2023; 150(1):dev201036. doi:10.1242/dev.201036.(This is a Cover Page article; andFeatured in the Journal’s Subject Collection)
- Manikandan M, Gadre S, Chhatar S, Chakrabarty G, Ahmed M, Patra C* and Patra M*.Potent Ruthenium-Ferrocene Bimetallic Antitumor Antiangiogenic Agent That Circumvents Platinum Resistance: From Synthesis and Mechanistic Studies to In Vivo Evaluation in Zebrafish.J Med Chem.(ISSN#0022-2623) 2022 Dec 22;65(24):16353-16371. doi: 10.1021/acs.jmedchem.2c01174.
- Satarkar D, and Patra C*. Evolution, Expression and Functional Analysis of CXCR3 in Neuronal and Cardiovascular Diseases: A Narrative Review. Front Cell Dev Biol. (ISSN#2296-634X)2022 Jun 20;10:882017. doi: 10.3389/fcell.2022.882017.
- Joshi B, Gaur H, Hui SP, and Patra C*. Celsr family genes are dynamically expressed in embryonic and juvenile zebrafish. Developmental Neurobiology (ISSN#1932-8451)2022 Mar;82(2):192-213.doi: 10.1002/dneu.22868.
- Joshi B, Wagh G, Kaur H, and Patra C*.Zebrafish Model to Study Angiotensin II-Mediated Pathophysiology. Biology(ISSN#2079-7737) 2021; 10(11), 1177.https://doi.org/10.3390/biology10111177
- Mukherjee D, Wagh G, Mokalled MH, Kontarakis Z, Dickson AL, Rayrikar A, Günther S, Poss KD, Stainier DYRand Patra C*.Ccn2a/Ctgfa is an injury-induced matricellular factor that promotes cardiac regeneration in zebrafish. Development(ISSN#0950-1991) 2021; 148, dev193219. doi:10.1242/dev.193219.
- .Pandey S, Mukherjee D, Kshirsagar P, Patra Cand Bodas D.Multiplexed bio-imaging using cadmium telluride quantum dots synthesized by mathematically derived process parameters in a continuous flow active microreactor.Materials Today Bio(ISSN#2590-0064) 2021; 11, 100123.
- Barrodia P, Patra C and Swain RK. EF-hand domain containing 2 (Efhc2) is crucial for distal segmentation of pronephros in zebrafish. Cell & Bioscience (ISSN#2045-3701) 2018; 8:53.
- Patra C*, Kontarakis Z, Kaur H, Rayrikar A, Mukherjee D and Stainier DYR. The zebrafish ventricle: A hub of cardiac endothelial cells for in vitro cell behavior studies. Scientific Reports (ISSN#2045-2322) 2017; 7:2687.
- Mokalled MH, Patra C, Dickson AL, Endo T, Stainier DYR and Poss KD. Injury-induced ctgfa directs glial bridging and spinal cord regeneration in zebrafish. Science (ISSN#1095-9203) 2016; 354 (6312):630-634.
- Patra C,Boccaccini AR and Engel FB. Vascularization for cardiac tissue engineering: the extracellular matrix. Thrombosis and Haemostasis (ISSN#1538-7836) 2015; 113(3):532-47. a: Contributed equally.
* Corresponding Author.